This blog post was originally published at NVIDIA’s website. It is reprinted here with the permission of NVIDIA.
The most commonly diagnosed cancer in the US today is skin cancer. There are three main variants: melanoma, basal cell carcinoma (BCC), and squamous cell carcinoma (SCC). Though melanoma only accounts for roughly 1% of all skin cancers, it is the most fatal, metastasizing rapidly without early detection and treatment. This makes early detection critical, as numerous studies show significantly better survival rates when detection is done in its earliest stages.
The current diagnosis procedure is done through a visual examination by a dermatologist, followed by a biopsy to confirm any suspected pathology. This manual examination is dependent on human subjectivity and thus suffers from error at a concerning rate. When a primary care physician looks for skin cancer, their sensitivity, or ability to identify a patient with the disease correctly, is only 0.45, while a dermatologist has a sensitivity of 0.97.
In recent years, the use of deep learning to perform medical diagnostics has become a quickly growing field. In this post, we discuss developing an end-to-end example of how deep learning could lead to an automated dermatology exam system free of human bias, using the recently announced NVIDIA Clara AGX development kit.
Datasets and models
This reference application is the pairing of two deep learning models:
- An object detection model (YOLOv4) that looks for moles on the body through a camera. This model was trained with an original dataset created from annotating body mole images.
- A classification model (EfficientNet) that receives moles from the object detection model and then determines if it is benign, unknown, or melanoma. The classification model was trained using the SIIM-ISIC melanoma Kaggle challenge dataset.
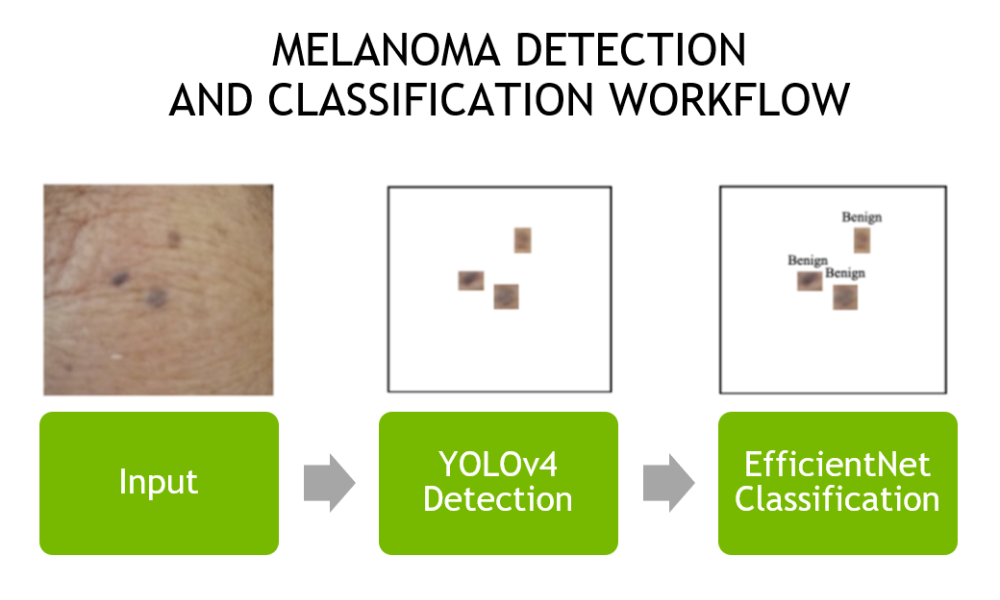
Figure 1 shows the workflow of the algorithm using a single video frame. The application can use a high-definition webcam or IP camera as input to the models, or even run on a previously captured video.
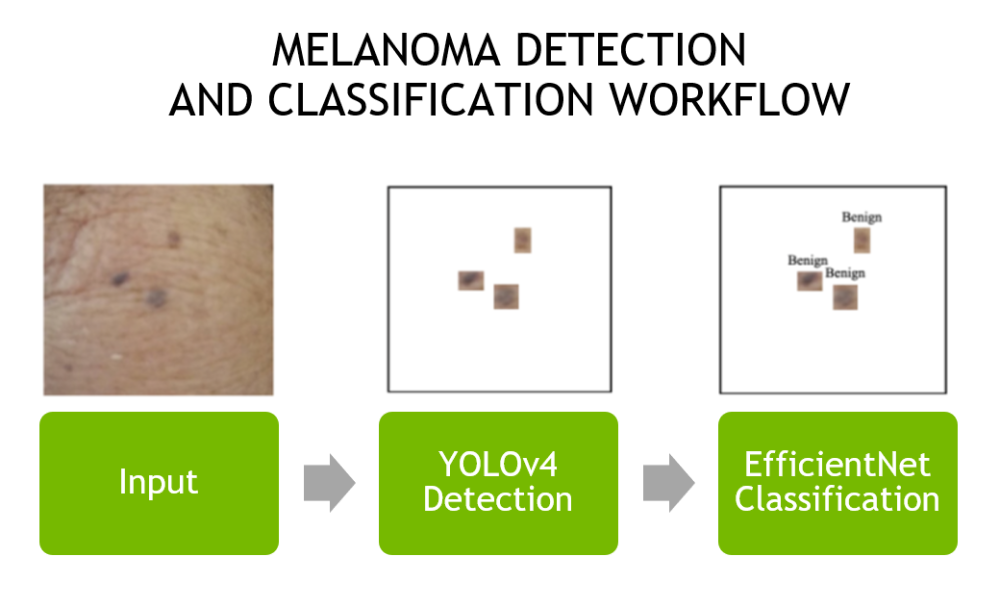
Figure 1. Skin mole detection and classification workflow.
Clara AGX development kit
This reference application was built using the NVIDIA Clara AGX development kit, a high-end performance workstation built with medical applications in mind. The system includes an RTX 6000 GPU, delivering 200+ INT8 AI TOPs of peak performance and 24 GB of VRAM, leaving plenty of overhead for running multiple models.
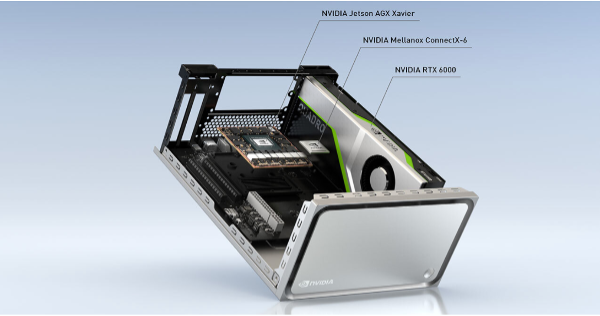

Figure 2. Clara AGX Developer Kit.
In addition, the AGX platform offers support for high bandwidth sensors through 100G Ethernet and an NVIDIA ConnectX-6 network interface card (NIC). NVIDIA partners are currently using the NVIDIA Clara AGX development kit to develop applications in ultrasound, genomics, and endoscopy.
The Clara AGX Developer Kit is currently available exclusively for members of the NVIDIA Clara Developer Partner Program. After you register, we’ll be in touch.
Summary
We’ve provided a research prototype of a dermatology application, but what would it take to transform this into a real application?
- Commercially usable data. The SIIM-ISIC dataset is strictly for non-commercial use.
- A much larger object detection dataset. The dataset that we used consisted of only a few hundred annotated images, which did lead to a larger than desired number of false positives.
- Run the models at the “speed of light” (SOL). SOL often entails training models to run using mixed precision and then transforming the models to work with the NVIDIA TensorRT framework. TensorRT is designed to optimize model inference on NVIDIA GPUs and work with common frameworks such as PyTorch and TensorFlow. These steps would help to ensure that your application pipeline runs in real-time.
- FDA clearance. Any developed medical application must be cleared by the FDA. Today, there are over 70 FDA-cleared AI applications, and the FDA has been active in soliciting feedback from developers in this area. This is typically a long (18 months) and arduous process, but a necessary one.
For more information, see the dermatology reference Docker container on NGC.
Roujia Wang
Ph.D. candidate, Duke University
Sean Huver, Ph.D.
Senior Machine Learning Engineer, Clara AGX Team, NVIDIA